Host-gene signals point to gut physiology as main driver of microbiome diversity
TL;DR Summary
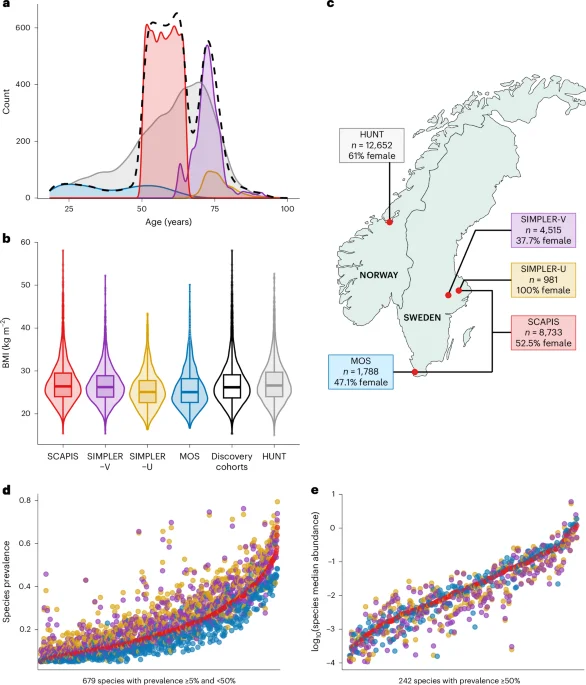
A large GWAS of harmonized gut metagenomic data from 16,017 Swedish adults with replication in 12,652 Norwegians identifies host genetic variants linked to microbiome richness and 149 species; the strongest signal at OR51E1–OR51E2 suggests enteroendocrine fatty acid sensing shapes microbial communities, with additional loci near mucin genes and bile-acid pathways. Mendelian randomization links some taxa to LDL cholesterol and BMI-related traits, underscoring gastrointestinal physiology as a key driver of microbiome variation. Limitations include European-ancestry focus and challenges in pinpointing causal genes.
Topics:health#enteroendocrine-cells#genome-wide-association-study#gut-microbiome#host-genetics#science#short-chain-fatty-acids
- Genome-wide association analyses highlight the role of the intestinal molecular environment in human gut microbiota variation Nature
- 11 genetic variants affect gut microbiome Uppsala universitet
- 11 Genetic Variants Linked to Gut Microbiome Composition via GWAS Inside Precision Medicine
- The HUNT study identifies host genetic factors reproducibly associated with human gut microbiota composition Nature
- Eleven Genetic Variants Influence the Gut Microbiome, New Study Reveals Bioengineer.org
Reading Insights
Total Reads
0
Unique Readers
8
Time Saved
67 min
vs 67 min read
Condensed
99%
13,397 → 81 words
Want the full story? Read the original article
Read on Nature