Structure-Powered Expansion of Eukaryote-Like Toolkit in Asgard Archaea
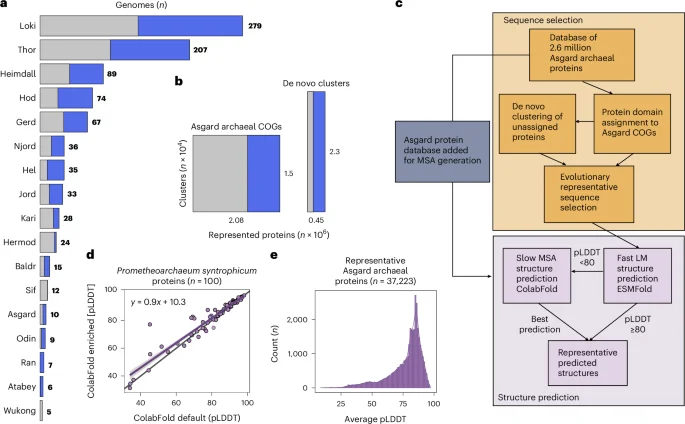
A large-scale structural analysis of 936 Asgard archaeal genomes using ColabFold and protein language models identified 1,319 novel isomorphic ESPs (iESPs) and organized them into 908 structural clusters, greatly expanding the repertoire of eukaryotic-like proteins in Asgard archaea. The iESPs are enriched in information storage/processing and cellular signaling, including components related to the vault (MVP) and Commander (COMMD) complexes, suggesting a higher degree of eukaryote-like cellular complexity in the archaeal ancestor of eukaryotes. The study also reveals patchy distributions and emphasizes that structural similarity, not just sequence similarity, drives annotation, highlighting the value of structure-based approaches for tracing deep evolutionary connections. The findings support a more nuanced view of eukaryogenesis and provide a methodological blueprint for uncovering distant homologues in non-model organisms.
- Prediction of eukaryotic cellular complexity in Asgard archaea using structural modelling Nature
- Mysterious Asgard microbes may point to origins of complex life CNN
- Ancient Microbes Fused Into Earth’s First Complex Cells. Scientists Just Learned What Made It Possible. Popular Mechanics
- Unveiling Eukaryotic Complexity in Asgard Archaea Structures BIOENGINEER.ORG
- Microbial Ancestor of Complex Life More Sophisticated Mirage News
Reading Insights
1
1
56 min
vs 57 min read
99%
11,387 → 122 words
Want the full story? Read the original article
Read on Nature